Software
Positively Split Ewald (PSE) Algorithm for RPY Hydrodynamics
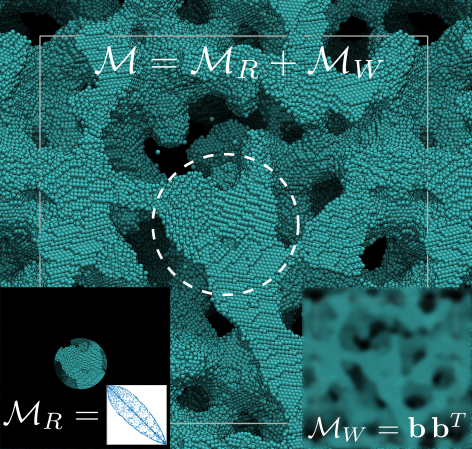 In the article "Rapid sampling of stochastic displacements in Brownian dynamics simulations", Fiore et al. present a new technique for sampling the stochastic thermal displacements in Brownian Dynamics simulations with hydrodynamic interactions represented at the Rotne-Prager-Yamawaka (RPY) level of approximation. This technique, called Positively-Split Ewald sampling, samples the Brownian displacements from a superposition of two indepedent distributions: a wave space (far-field or long-ranged) contribution, computed using techniques from fluctuating hydrodynamics and non-uniform fast Fourier transforms; and a real space (near-field or short-ranged) correction, computed using a Krylov subspace method. The total computational complexity of the algorithm scales linearly with the number of particles modeled. The PSE algorithm enables simulations of up to 4 million colloidal particles. A high-performance implementation of the PSE algorithm on graphics processing units (GPUs), written as a plugin for the software package HOOMD-blue is freely available on GitHub.
In the article "Rapid sampling of stochastic displacements in Brownian dynamics simulations", Fiore et al. present a new technique for sampling the stochastic thermal displacements in Brownian Dynamics simulations with hydrodynamic interactions represented at the Rotne-Prager-Yamawaka (RPY) level of approximation. This technique, called Positively-Split Ewald sampling, samples the Brownian displacements from a superposition of two indepedent distributions: a wave space (far-field or long-ranged) contribution, computed using techniques from fluctuating hydrodynamics and non-uniform fast Fourier transforms; and a real space (near-field or short-ranged) correction, computed using a Krylov subspace method. The total computational complexity of the algorithm scales linearly with the number of particles modeled. The PSE algorithm enables simulations of up to 4 million colloidal particles. A high-performance implementation of the PSE algorithm on graphics processing units (GPUs), written as a plugin for the software package HOOMD-blue is freely available on GitHub.
Brownian Dynamics of Rigid Bodies
 A GPU algorithm for Brownian Dynamics of composite-bead rigid particles is implemented. Arbitrarily shaped colloidal particles or macromolecules can be modeled as rigid assemblies of beads from surface tessellation. Hydrodynamic interactions among the beads are modeled with RPY tensor applying the PSE algorithm. Correct hydrodynamic and transport properties are successfully captured as demonstrated in the article "Rapid calculation of hydrodynamic and transport properties in concentrated solutions of colloidal particles and macromolecules". The software package can be downloaded here.
A GPU algorithm for Brownian Dynamics of composite-bead rigid particles is implemented. Arbitrarily shaped colloidal particles or macromolecules can be modeled as rigid assemblies of beads from surface tessellation. Hydrodynamic interactions among the beads are modeled with RPY tensor applying the PSE algorithm. Correct hydrodynamic and transport properties are successfully captured as demonstrated in the article "Rapid calculation of hydrodynamic and transport properties in concentrated solutions of colloidal particles and macromolecules". The software package can be downloaded here.
Teaching Stokesian Dynamics to Swim
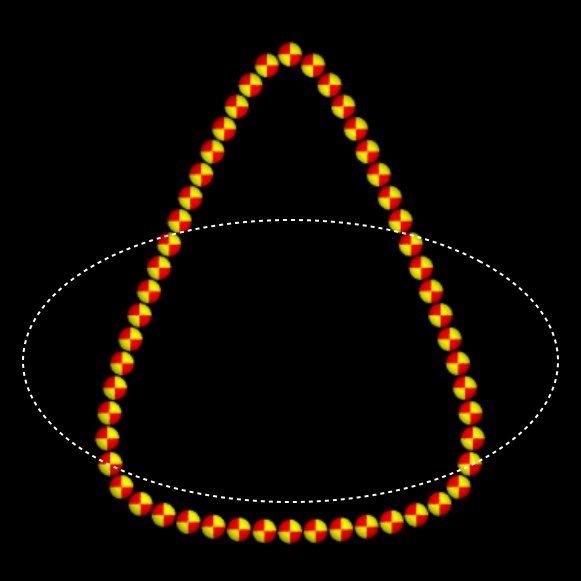 In the article "Modeling hydrodynamic self-propulsion with Stokesian Dynamics.
Or teaching Stokesian Dynamics to swim," Swan, Brady and Moore demonstrate how models of swimming microorganisms can
be constructed from rigid particles that move along a prescribed gait. The hydrodynamic interactions between particles
serve as a surrogate for a solution of the Stokes equations dictating the fluid flow around the swimmer. These interactions
are accounted for faithfully with the Stokesian Dynamics method. Click
this link to view some movies of swimmers modeled in the article. A software
package capable replicating these model swimmers as well as others is available.
In the article "Modeling hydrodynamic self-propulsion with Stokesian Dynamics.
Or teaching Stokesian Dynamics to swim," Swan, Brady and Moore demonstrate how models of swimming microorganisms can
be constructed from rigid particles that move along a prescribed gait. The hydrodynamic interactions between particles
serve as a surrogate for a solution of the Stokes equations dictating the fluid flow around the swimmer. These interactions
are accounted for faithfully with the Stokesian Dynamics method. Click
this link to view some movies of swimmers modeled in the article. A software
package capable replicating these model swimmers as well as others is available.
Force spectroscopy near interfaces
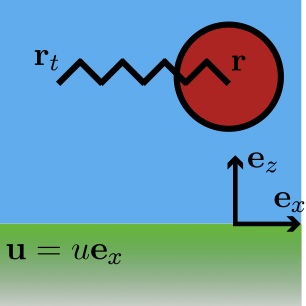 A common practice in soft matter physics is to use colloidal probes to measure inter- and intra-molecular forces. For instance,
A colloidal bead may be tethered to a microscope slide with DNA. When a force is applied to the bead, the DNA is stretched and
a force-extension curve can be inferred. The key then is to know the force exerted on the bead. One method of "pulling" on the bead
is to generate a flow field which drags it along the slide. When far from the surface of the slide, the drag coefficient of the bead
is well know. In the article "Nonequilibrium distributions and hydrodynamic coupling distort the measurement of nanoscale forces near interfaces,"
Swan and Furst show that when the bead is near the interface, its Brownian motion makes knowledge of its drag coefficient difficult.
They derive a new model for the force-extension curve associated with a bead tethered by a Hookean spring that accounts for the
this stochastic motion explicitly. A software package capable of replicating the results of this model is available.
A common practice in soft matter physics is to use colloidal probes to measure inter- and intra-molecular forces. For instance,
A colloidal bead may be tethered to a microscope slide with DNA. When a force is applied to the bead, the DNA is stretched and
a force-extension curve can be inferred. The key then is to know the force exerted on the bead. One method of "pulling" on the bead
is to generate a flow field which drags it along the slide. When far from the surface of the slide, the drag coefficient of the bead
is well know. In the article "Nonequilibrium distributions and hydrodynamic coupling distort the measurement of nanoscale forces near interfaces,"
Swan and Furst show that when the bead is near the interface, its Brownian motion makes knowledge of its drag coefficient difficult.
They derive a new model for the force-extension curve associated with a bead tethered by a Hookean spring that accounts for the
this stochastic motion explicitly. A software package capable of replicating the results of this model is available.
Brownian Dynamics, LIVE
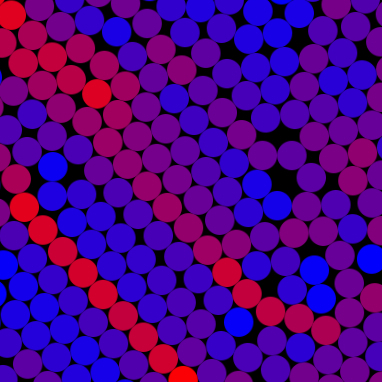 Brownian dynamics is a technique for modeling a dispersion small particles and macromolecules on time scales long relative the inertial
relaxation time of the fluid. Hydrodynamic interactions between the particles are neglected, but the Brownian motion and
any conservative interactions are accurately accounted for. Such a model is particularly useful when the constituents of
the dispersion are highly repulsive. Within this page, a Brownian dynamics simulation is
executed live in the browser window using HTML5 and javascript. The particles are colored to indicate the stress exerted on their surface.
From such a simple model complex particle and stress structures emerge.
Brownian dynamics is a technique for modeling a dispersion small particles and macromolecules on time scales long relative the inertial
relaxation time of the fluid. Hydrodynamic interactions between the particles are neglected, but the Brownian motion and
any conservative interactions are accurately accounted for. Such a model is particularly useful when the constituents of
the dispersion are highly repulsive. Within this page, a Brownian dynamics simulation is
executed live in the browser window using HTML5 and javascript. The particles are colored to indicate the stress exerted on their surface.
From such a simple model complex particle and stress structures emerge.
Micro-rheology, LIVE
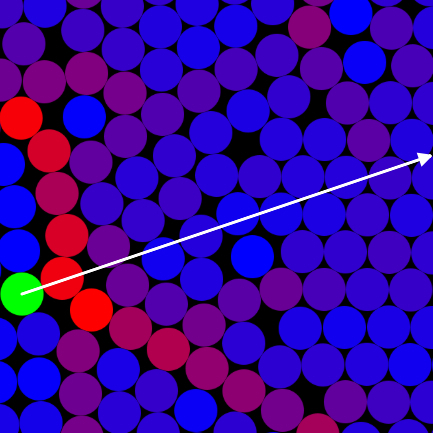 Micro-rheology is a technique for interrogating the visco-elastic properties of complex fluids by analyzing the motion
of colloidal probes embedded within. In passive micro-rheology, a colloidal probe moves solely due to the Brownian force
and its mean squared displacement related to the creep compliance of the complex fluid via the generalized Stokes-Einstein-Sutherland
relation. In active micro-rheology, a colloidal probe is driven through the complex fluid by an external force and
the viscosity of the fluid is inferred from the probes velocity via the Stokes drag law. This,
simulates the active micro-rheology of a colloidal dispersion live in the browser window using HTML5 and javascript.
Micro-rheology is a technique for interrogating the visco-elastic properties of complex fluids by analyzing the motion
of colloidal probes embedded within. In passive micro-rheology, a colloidal probe moves solely due to the Brownian force
and its mean squared displacement related to the creep compliance of the complex fluid via the generalized Stokes-Einstein-Sutherland
relation. In active micro-rheology, a colloidal probe is driven through the complex fluid by an external force and
the viscosity of the fluid is inferred from the probes velocity via the Stokes drag law. This,
simulates the active micro-rheology of a colloidal dispersion live in the browser window using HTML5 and javascript.
Stokesian Dynamics, LIVE
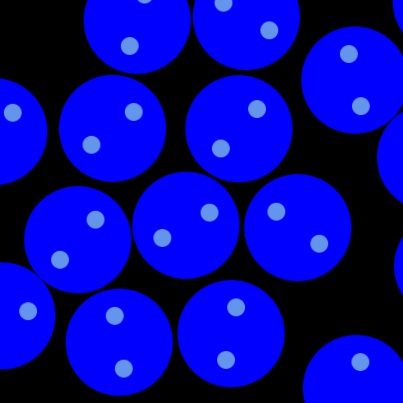 Stokesian Dynamics is a technique for modeling the low Reynolds number hydrodynamic interaction between microscopic particles.
When micron sized particles are immersed in a viscous fluid, the diffusion of vorticity is practically instantaneous. Thus,
fluid mechanics are greatly simplified. Particles separated by large distances interact through the fluid via a "1-over-r"
force law. The relative motion between nearly touching particles can give rise to incredibly strong "lubrication" forces.
The Stokesian Dynamics method is designed to accurately account for both the long-ranged many body forces and the short-ranged
lubrication forces among thousands of suspended particles. Here is a game built in HTML5
and javascript in which hydrodynamically interacting particles must be dragged and then dropped within target regions. The hydrodynamic
interactions are calculated using the Stokesian Dynamics method.
Stokesian Dynamics is a technique for modeling the low Reynolds number hydrodynamic interaction between microscopic particles.
When micron sized particles are immersed in a viscous fluid, the diffusion of vorticity is practically instantaneous. Thus,
fluid mechanics are greatly simplified. Particles separated by large distances interact through the fluid via a "1-over-r"
force law. The relative motion between nearly touching particles can give rise to incredibly strong "lubrication" forces.
The Stokesian Dynamics method is designed to accurately account for both the long-ranged many body forces and the short-ranged
lubrication forces among thousands of suspended particles. Here is a game built in HTML5
and javascript in which hydrodynamically interacting particles must be dragged and then dropped within target regions. The hydrodynamic
interactions are calculated using the Stokesian Dynamics method.