|

Greg Randall
Ph.D Candidate
Chemical Engineering, MIT
gregr[at]email.unc.edu
|
Greg Randall
Education:
B.S. Chemical Engineering, Cornell University, 1998
Research Interests:
Single Molecule Analysis of DNA in Electric Fields
G. Randall and P. Doyle
Sponsorship: National Science Foundation Grant CTS-0239012
Recent advances in gene therapy and crime investigation have spurred a
demand for rapid gene mapping of large (kbp-Mbp) DNA
molecules. Because current electrophoresis technologies are inadequate for large DNA, several promising MEMS designs for DNA
mapping have been recently proposed that require either 1) a DNA molecule negotiating an obstacle course in a microchannel or 2)
stretching a DNA coil for linear analysis. The goal of our research is to experimentally probe the fundamental physics that
underlie these DNA mapping designs. In general, the governing physics is complex due to the confinement of the microchannel, the
coiled-nature of long DNA molecules, and the induced electric field gradients from obstacles and changes in channel dimensions.
With single molecule microscopy, we have demonstrated many of the governing physical mechanisms at play in these gene mapping
microfluidic devices [1-3]. For example, we have shown the experimental scaling for the diffusion coefficient of DNA in a confined
channel (Figure 1a) and the probability distribution for the collision time of a DNA molecule unhooking from a small obstacle
(Figure 1c). In addition, we have thoroughly investigated DNA stretching in electric field gradients created by a contraction and
an obstacle (Figure 2). Just as a flow gradient stretches a polymer, an electric field gradient can stretch a charged polymer like
DNA. Because electric field gradients have no local rotational components, a charged polymer will experience purely extensional
deformation. These findings will aid the design of DNA separation devices that contain many obstacles and contractions and they
also offer an attractive way to completely stretch DNA for linear analysis.
|
|
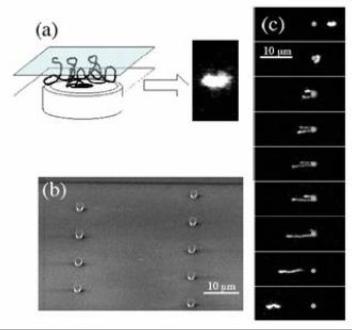 |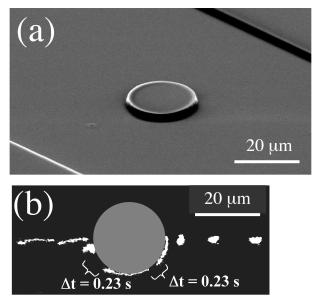  |
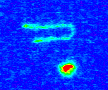
Figure 1: (a) Cartoon of a long DNA molecule in a thin slit over a microscope objective and a sample experimental image.
(b) SEM image of a disperse array of small PDMS (polydimethylsiloxane) obstacles (Robs=0.8 μm,
height=2 μm). (c) A
hooking collision of λ-DNA with one of the small obstacles (0.17 s intervals, DNA moving right to left).
|
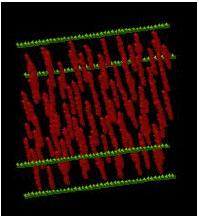
Figure 2: (a) SEM image of a large PDMS obstacle (Robs=10 μm, height=2 μm).
(b) A center-line collision of DNA with the large obstacle (0.33 s intervals unless noted, DNA moving right to left).
|
|
|
|
|
References:
[1] G. C. Randall and P. S. Doyle, "Electrophoretic collision of a DNA molecule with an insulating post," Phys. Rev.
Lett., 93, 058102, (2004).
[2] Y.-L. Chen, M. D. Graham, J. J. dePablo, G. C. Randall, M. Gupta, and P. S. Doyle, "Conformation and dynamics of
single DNA in parallel-plate slit microchannels", Phys. Rev. E, 70, 060901, (2004).
[3] G. C. Randall and P. S. Doyle, "DNA deformation in electric fields: DNA driven past a cylindrical obstruction",
Macromolecules, 38, 2410-2418, (2005).
|
|
|